by Andrea Navas-Olive, Enrique R Sebastian & Liset M de la Prida
In many occasions you may need to classify electrophysiological events (e.g. ripples, theta cycles, up-down states) in an unbiased manner, just relaying in the variability of their waveforms. One way to approach this problem is to represent each event as a point in the high-dimensional space that results from the recording sampling rate. In such a space, similar events will cloud together. The problem is how to visualize them all.
This can be done using Self-Organizing Maps (SOM). SOM is an artifical neural network that learn to generate a low dimensional representation of a manifold (i.e. the cloud) in a higher dimensional space. Typically, SOM is used to reduce dimensionality to the plane. To put it simple, SOM learns how to better cover the cloud in the high-dimensional space with a sheet (i.e. a bidimensional matrix) so that we can see how the dots are scattered in the sheet. SOM looks to preserve the properties of the cloud while imprinting it in the sheet.
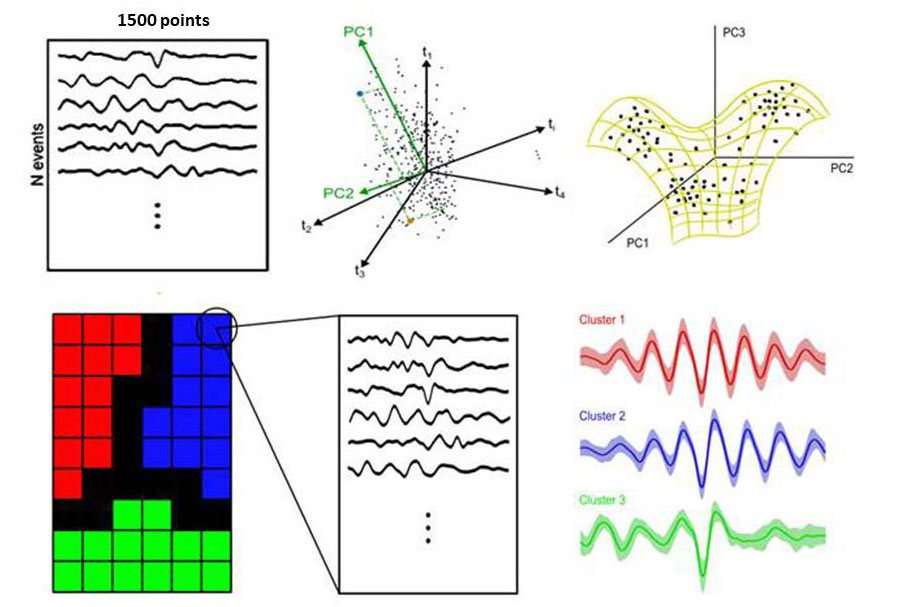
For instance, imagine you have N events sampled as 1500 time points. Each event will be a dot in the 1500-d space. You can directly try to fit the sheet with SOM in that space, which probably requires lot of computer power, or you can apply some intermediate reduction to capture the axis of major variations using principal component analysis (PCA) first. Whatever you decide, you can then use SOM to fit the sheet either in the 1500-d space or in the PCA-d space. SOM looks for stretching the sheet to fit in the cloud and so you end up with your events being differently distributed according to their similarity (e.g. as cluster 1, 2 and 3 in the example).
We developed an application to perform this analysis. RhythSOM is a computational tool for MATLAB to classify brain rhythms or events, such hippocampal sharp wave ripples or theta cycles, in a unsupervised way, just by their waveform.
RhythSOM Github: https://github.com/acnavasolive/RhythSOM
See Valero et al., Neuron 94(6), 1234-1247 (2017) for application in sharp-wave ripples and Navas-Olive et al, Nat Commu 11, 2217 (2020) for application in theta cycles